Diagnosis of autism from biomarkers is a holy grail for
biomedical researchers. The days when it was thought we would find “the autism
gene” are long gone, and it’s clear that both the biology and the psychology of
autism is highly complex and heterogeneous. One approach is to search for
individual genes where mutations are more likely in those with autism. Another
is to address the complexity head-on by looking for combinations of biomarkers
that could predict who has autism. The
latter approach is adopted in a paper by Bao et al (2022) who claimed that an
ensemble of gene expression measures taken from blood samples could accurately
predict which toddlers were autistic (ASD) and which were typically-developing (TD). An anonymous commenter on PubPeer
queried whether the method was as robust as the authors claimed, arguing that
there was evidence for “overfitting”. I was asked for my thoughts by a
journalist, and they were complicated enough to merit a blogpost. The bottom line is that there are reasons to
be cautious about the conclusion of the authors that they have developed “an
innovative and accurate ASD gene expression classifier”.
Some of the points I raise here applied to a previous biomarker study that I blogged about in 2019. These are general issues about the mismatch between what is done in typical studies in this area and what is needed for a clinically useful screening test.
Base rates
Consider first how a screening test might be used. One possibility is that there might be a move towards universal screening, allowing early diagnosis that might help ensure intervention starts young. But for effective screening in that context, you need extremely high diagnostic accuracy, and accuracy depends on the frequency of autism in the population. I discussed this back in 2010. The levels of accurate classification reported by Bao et al would be of no use for population screening because there would be an extremely high rate of false positives, given that most children don’t have autism.
Diagnostic specificity
But, you may say, we aren’t talking about universal screening. The test might be particularly useful for those who either (a) already have an older child with autism, or (b) are concerned about their child’s development. Here the probability of a positive autism diagnosis is higher than in the general population. However, if that’s what we are interested in, then we need a different comparison group – not typically-developing toddlers, but unaffected siblings of children with autism, and/or children with other neurodevelopmental disorders.
When I had a look at the code that the authors deposited for data analysis, it implied that they did have data on children with more general developmental delays, and sibs of those with autism, but they are not reported in this paper.
The analyses done by the researchers are extremely complex and time-consuming, and it is understandable that they may prefer to start out with the clearest case of comparing autism with typically-developing children. But the acid test of the suitability of the classifier for clinical use would be a demonstration that it could distinguish children with autism from unaffected siblings, and from nonautistic children with intellectual disability.
Reliability of measures
If you run a diagnostic test, an obvious question is whether you’d get the same result on a second test run. With biological and psychological measures the answer is almost always no, but the key issue for a screener is just how much change there is. Gene expression levels could vary from occasion to occasion depending on time of day or what you’d eaten – I have no idea how important this might be, but it's not possible to evaluate in this paper, where measures come from a single blood sample. My personal view is that the whole field of biomedical research needs to wake up to the importance of reliability of measurement so that researchers don’t waste time exploring the predictive power of measures that may be too unreliable to be useful. Information about stability of measures over time is a basic requirement for any diagnostic measure.
A related issue concerns comparability of procedures for autism and TD groups. Were blood samples collected by the same clinicians over the same period and processed in the same lab for these two groups? Were the blood analyses automated and/or done blind? It’s crucial to be confident that minor differences in clinical or lab procedures do not bias results in this kind of study.
Overfitting
Overfitting is really just a polite way of saying that the data may be noise. If you run enough analyses, something is bound to look significant, just by chance. In the first step of the analysis, the researchers ran 42,840 models on “training” data from 93 autistic and 82 TD children and found 1,822 of them performed better than .80 on a measure that reflects diagnostic accuracy (AUC-ROC – which roughly corresponds to proportion correctly classified: .50 is chance, and 1.00 is perfect classification). So we can see that just over 4% of the models (1822/42840) performed this well.
The researchers were aware of the possibility of overfitting, and they addressed it head-on, saying: “To test this, we permuted the sample labels (i.e., ASD and TD) for all subjects in our Training set and ran the pipeline to test all feature engineering and classification methods. Importantly, we tested all 42,840 candidate models and found the median AUC-ROC score was 0.5101 with the 95th CI (0.42–0.65) on the randomized samples. As expected, only rare chance instances of good 'classification' occurred.” The distribution of scores is shown in Figure 2b.
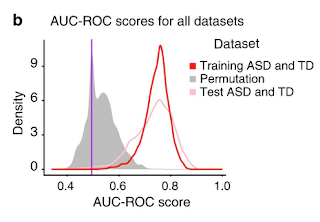 |
| Figure 2b from Bao et al (2022) |
They then ran a further analysis on a “test set” of 34 autistic and
31 TD children who had been held out of the original analysis, and found that
742 of the 1822 models performed better than .80 in classification. That’s 40%
of the tested models. Assuming I have
understood the methods correctly, that does look meaningful and hard to explain
just in terms of statistical noise. In
effect, they have run a replication study and found that a substantial subset
of the identified models do continue to separate autism and TD groups when new
children are considered. The claim is that there is substantial overlap in the models that fall in the right-hand area under the curve for the red and pink distributions.
The PubPeer commenter seems concerned that results look too good to be true. In particular, Figure 2b suggests the models perform a bit better in the test set than in the training set. But the figure shows the distribution of scores for all the models (not just the selected models) and, given the small sample sizes, the differences between distributions does not seem large to me. I was more surprised by the relatively tight distribution of AUC-ROC values obtained in the permutation analysis, as I would have anticipated some models would have given high classification accuracy just by chance in a sample of this size.
The researchers went on to present data for the set of models that achieved .8 classification in both training and test sets. This seemed a reasonable approach to me. The PubPeer commenter is correct in arguing that there will be some bias caused by selecting models this way, and that one would expect less good performance in a completely new sample, but the 2-stage selection of models would seem to ensure there is not "massive overfitting". I think there would be a problem if only 4% of the 1822 selected models had given accurate classification, but the good rate of agreement between the models selected in the training and test samples, coupled with the lack of good models in the permuted data, suggests there is a genuine effect here.
Conclusion
So, in sum, I think that the results can’t just be attributed to overfitting, but I nevertheless have reservations about whether they would be useful for screening for autism. And one of the first things I’d check if I were the researchers would be the reliability of the diagnostic classification in repeated blood samples taken on different occasions, as that would need to be high for the test to be of clinical use.
Note: I'd welcome comments or corrections on this post. Please note, comments are moderated to avoid spam, and so may not appear immediately. If you post a comment and it has not appeared in 24 hr, please email me and I'll ensure it gets posted.
PS. See comment from original PubPeer poster attached.
Also, 8th Dec 2022, I added a further PubPeer comment asking authors to comment on Figure 2B, which does seem odd.
https://pubpeer.com/publications/B693366B2B51D143C713359F151F7B#4
PPS. 10th Dec 2022.
Author Eric Courchesne has responded to several of the points made in this blogpost on Pubpeer: https://pubpeer.com/publications/B693366B2B51D143C713359F151F7B#5












I'm the anonymous pubpeer.
ReplyDeleteFirst a minor correction: A correct description of the interpretation of the AUROC would be "the probability that, given a random positive and negative case, the classifier would give a higher score to the positive” (where by ‘score’ I mean the output of the classifier and what is used to make the ranking for the curve). Not “the proportion correctly classified”. “How well the classifier is working” would have been fine for a lay audience.
A more important correction is that “overfitting” isn’t synonymous with “your data may be noise” (politely or otherwise). Here it means that “the classifier is too specific to the data on hand and might not generalize to new data”. This can happen even when there is a very strong signal in the data at hand. I acknowledge that the term “overfitting” can be used a bit vaguely but “noise” is not directly implied by overfitting.
Failure to generalize can still happen even without (blatant) overfitting, but overfitting in the way that the authors did makes a failure to generalize a much greater risk.
I do not doubt that the authors’ classifier works with the stated performance on their data, and they are not just picking up “noise” (at least, that’s a separate issue). The concern is that when this is tried on new data, performance will decrease. As far as I can see, their experimental design does not address this because there is no “new data” that wasn’t used in developing the final classifier.
The performance on the “held out” 65 samples is much worse unless one has the “foresight” to cherry-pick a model that does well. And that’s what the authors did as far as I can tell. This is not up to the standards of the field.
Failure to generalize has been the downfall of many (most?) biomarker searches for brain conditions. Things look great, patents are filed, companies are started, investors come. Where are all those biomarkers now?
We can quibble about the risk of “massive” overfitting vs. “one would expect less good performance in a completely new sample” (“less good” is just a polite way of saying “worse”?).
My point is that the issue could have been avoided (at least provisionally) if the authors had put some of their data in “escrow” to be tested on after all was said and done, and this well-understood best practice. The usual excuse is, “We wouldn’t have enough data if we did it the right way!”. Well, too bad. Science, and the autism community, deserves better.